Abraham J. Koo

Associate Professor
Department of Biochemisty
E-mail: kooaj at missouri dot edu
Office address: 127 Schweitzer Hall
Office phone: 573-882-9227
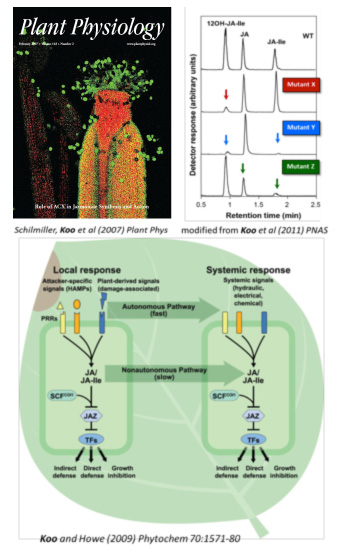 Our lab’s research involves the study of small signaling molecules that cells use to detect extracellular stimuli and to coordinate intra- and intercellular responses. To compensate for their lack of mobility, plant cells rely heavily on chemical cues to interact with their surroundings—one reason why plants are so rich in pharmaceuticals and provide attractive models for studying chemically mediated cell signaling. We use highly sensitive and selective mass-spectrometry-based methodologies to capture and to monitor plants’ chemical messengers, which often exist in trace amounts. Using powerful analytical tools combined with the rich genetic and genomic resources available for the model plant Arabidopsis has allowed us to identify new players in defense signaling cascades within a plant’s immune response against insects and fungal pathogens. Recently, our focus has been to describe mechanisms that control optimal levels of plant hormone jasmonates (JAs). JAs are a group of oxylipins produced by the oxidative metabolism of polyunsaturated fatty acids, analogous to eicosanoids in animals. Similar to eicosanoids, JAs play an important role in mechanical stress-induced responses and plant immunity. Thus, knowledge obtained from this research has implications for use in applied agronomy as well as for the advancement of foundational science in the area of cross-kingdom, conserved chemical cell signaling. In addition, because JAs have been reported to have anti-cancerous effects on certain tumor cells and are master regulators of secondary metabolites in plants, many of which are antioxidants beneficial to humans, the study of JAs will contribute to overall human health.
Our lab’s research involves the study of small signaling molecules that cells use to detect extracellular stimuli and to coordinate intra- and intercellular responses. To compensate for their lack of mobility, plant cells rely heavily on chemical cues to interact with their surroundings—one reason why plants are so rich in pharmaceuticals and provide attractive models for studying chemically mediated cell signaling. We use highly sensitive and selective mass-spectrometry-based methodologies to capture and to monitor plants’ chemical messengers, which often exist in trace amounts. Using powerful analytical tools combined with the rich genetic and genomic resources available for the model plant Arabidopsis has allowed us to identify new players in defense signaling cascades within a plant’s immune response against insects and fungal pathogens. Recently, our focus has been to describe mechanisms that control optimal levels of plant hormone jasmonates (JAs). JAs are a group of oxylipins produced by the oxidative metabolism of polyunsaturated fatty acids, analogous to eicosanoids in animals. Similar to eicosanoids, JAs play an important role in mechanical stress-induced responses and plant immunity. Thus, knowledge obtained from this research has implications for use in applied agronomy as well as for the advancement of foundational science in the area of cross-kingdom, conserved chemical cell signaling. In addition, because JAs have been reported to have anti-cancerous effects on certain tumor cells and are master regulators of secondary metabolites in plants, many of which are antioxidants beneficial to humans, the study of JAs will contribute to overall human health.
Kimberlin A, Holtsclaw RE, Zhang T, Mulaudzi T, and Koo AJ. (2022) On the initiation of jasmonate biosynthesis in wounded leaves. Plant Physiol. Apr 11:kiac163. doi: 10.1093/plphys/kiac163. Online ahead of print. PMID: 35404431
Wang M, Garneau M, Poudel AN, Lamm D, Koo AJ, Bates P, Thelen J. (2022) Overexpression of pea α-carboxyltransferase in Arabidopsis and Camelina increases fatty acid synthesis leading to improved seed oil content. Plant J. 110:1035-1046. doi: 10.1111/tpj.15721. PMID: 35220631
Kimberlin A, Holtsclaw RE, and Koo AJ. (2021) Differential regulation of the ribosomal association of mRNA transcripts in an Arabidopsis mutant defective in jasmonate-dependent wound response. Front. Plant Sci. 12:637959. doi: 10.3389/fpls.2021.637959. [PubMed]
Fernández-Milmanda GL, Crocco CD, Reichelt M, Mazza CA, Köllner TG, Zhang T, Cargnel MD, Lichy MZ, Koo AJ., Austin AT, Gershenzon J, and Ballaré CL. (2020) A light-dependent molecular link between competition cues 1 and defense responses in plants. Nature Plant 6:223-230. doi: 10.1038/s41477-020-0604-8. [PubMed]
Body MJ, Dave DF, Coffman CM, Paret TY, Koo AJ., Cocroft RB, and Appel HM. (2019) Plant response to insect feeding vibrations involves oxylipin signaling. Front. Plant Sci. 10. doi: 10.3389/fpls.2019.01586. [PubMed]
Poudel AN, Holtsclaw RE, Kimberlin A, Sen S, Zeng S, Joshi T, Lei Z, Sumner LW, Singh K, Matsuura H, and Koo AJ.. (2019) 12-Hydroxy-jasmonoyl-L-isoleucine is an active jasmonate that signals through CORONATINE INSENSITIVE 1 and contributes to the wound response in Arabidopsis. Plant Cell Physiol. 60: 2152-2166. doi: 10.1093/pcp/pcz109. [PubMed]
Lunde C, Kimberlin A, Leiboff S, Koo AJ., Hake S. (2019). Tasselseed5 overexpresses a wound-inducible enzyme, ZmCYP94B1, that affects jasmonate catabolism, sex determination, and plant architecture in maize. Commun Biol. 2:114. doi: 10.1038/s42003-019-0354-1. eCollection 2019. [PubMed]
Toyota M, Spencer D, Sawai-Toyota S, Jiaqi W, Zhang T, Koo AJ., Howe GA, Gilroy S. (2018). Glutamate triggers long-distance, calcium-based plant defense signaling. Science. 361(6407):1112-1115. doi: 10.1126/science.aat7744. [PubMed]
Yurchenko O, Kimberlin A, Mehling M, Koo AJ., Chapman KD, Mullen RT, Dyer JM. (2018). Response of high leaf-oil Arabidopsis thaliana plant lines to biotic or abiotic stress. Plant Signal Behav. 13(5):e1464361. doi: 10.1080/15592324.2018.1464361. [PubMed]
Howe GA, Major IT, Koo AJ.. (2018). Modularity in Jasmonate Signaling for Multistress Resilience. Annu Rev Plant Biol. 69:387-415. doi: 10.1146/annurev-arplant-042817-040047. [PubMed]
Tripathi D, Zhang T, Koo AJ., Stacey G, Tanaka K. (2018). Extracellular ATP Acts on Jasmonate Signaling to Reinforce Plant Defense. Plant Physiol. 176(1):511-523. doi: 10.1104/pp.17.01477. [PubMed]
Poudel AN, Zhang T, Kwasniewski M, Nakabayashi R, Saito K, Koo AJ.. (2016). Mutations in jasmonoyl-L-isoleucine-12-hydroxylases suppress multiple JA-dependent wound responses in Arabidopsis thaliana. Biochim Biophys Acta. 1861(9 Pt B):1396-1408. doi: 10.1016/j.bbalip.2016.03.006. [PubMed]
Schneider A, Aghamirzaie D, Elmarakeby H, Poudel AN, Koo AJ., Heath LS, Grene R, Collakova E. (2016). Potential targets of VIVIPAROUS1/ABI3-LIKE1 (VAL1) repression in developing Arabidopsis thaliana embryos. Plant J. 85(2):305-19. doi: 10.1111/tpj.13106. [PubMed]
Zhang T, Poudel AN, Jewell JB, Kitaoka N, Staswick P, Matsuura H, Koo AJ. (2016). Hormone crosstalk in wound stress response: wound-inducible amidohydrolases can simultaneously regulate jasmonate and auxin homeostasis in Arabidopsis thaliana. J Exp Bot. 67(7):2107-20. doi: 10.1093/jxb/erv521. [PubMed]
Nguyen PD, Pike S, Wang J, Nepal Poudel A, Heinz R, Schultz JC, Koo AJ., Mitchum MG, Appel HM, Gassmann W. (2016). The Arabidopsis immune regulator SRFR1 dampens defences against herbivory by Spodoptera exigua and parasitism by Heterodera schachtii. Mol Plant Pathol. 17(4):588-600. doi: 10.1111/mpp.12304. [PubMed]
Smith JM, Leslie ME, Robinson SJ, Korasick DA, Zhang T, Backues SK, Cornish PV, Koo AJ., Bednarek SY, Heese A. (2014). Loss of Arabidopsis thaliana Dynamin-Related Protein 2B reveals separation of innate immune signaling pathways. PLoS Pathog. 10(12):e1004578. doi: 10.1371/journal.ppat.1004578. eCollection 2014 Dec. [PubMed]
Koo AJ., Thireault C, Zemelis S, Poudel AN, Zhang T, Kitaoka N, Brandizzi F, Matsuura H, Howe GA. (2014). Endoplasmic reticulum-associated inactivation of the hormone jasmonoyl-L-isoleucine by multiple members of the cytochrome P450 94 family in Arabidopsis. J Biol Chem. 289(43):29728-38. doi: 10.1074/jbc.M114.603084. [PubMed]