Henry Nguyen

Curators' Professor
Division of Plant Science and Technology
E-mail: nguyenhenry at missouri dot edu
Web site: Soybean Genetics and Genomics Laboratory
Office address: 271d Bond Life Sciences Center
Office phone: 573-882-5494
Lab address: 25 Agriculture Building
Lab phone: 573-882-5483

Research in Dr. Nguyen’s laboratory is focused on soybean genomics and comparative genetics of plant stress responses. Three overall research areas are: (a) soybean genome mapping and molecular breeding, (b) functional genomics of drought tolerance, and (c) nutritional genomics.
Soybean cDNA array
Soybean genome mapping and molecular breeding: Our laboratory is part of a national effort aimed at the development of a high resolution physical map of William 82 soybean genome based on restriction fingerprinting of BAC clones. We developed 6-dimensional BAC pools and contribute to the anchoring of genetic markers to the physical map. We are constructing a standard genetic map using close to 800 progenies from a cross of Forrest and Williams 82. Currently we focus on mapping QTL controlling SCN resistance and analysis of candidate genes for the SCN resistance response. Populations have been developed for mapping QTL and candidate gene analysis for isoflavone and sapponin accumulation in soybean. Our long range goal is to understand the genetic basis of QTL underlying important traits utilizing information from the integrated genetic and physical maps and to develop high throughput marker-assisted selection for soybean improvement in collaboration with the breeders.

Functional genomics of drought tolerance: Drought is a major abiotic stress limiting soybean productivity. Drought also affects soybean seed composition. A series of molecular, cellular and physiological changes occur in drought stressed plants. To understand the regulatory networks and mechanisms of drought tolerance, we use both forward and reverse genetic approaches. We currently characterize gene expression products at the transcriptome, proteome and metabolome levels. Using microarray, we are conducting gene expression profiling of soybean leaf and root under drought stress to elucidate the pattern of gene expression and signaling cascades. Laser capture micro-dissection technique is being used to target specific cell types. We validate expression results through q-RT PCR and gene function analysis. We are also conducting a comparative genomics study between soybean and model plants, Medicago truncatula and Arabidopsis under drought stress. We have sequenced and deposited more than 17,000 ESTs associated with root hair and drought responses in soybean roots. To support the transcript profile data, we are searching protein profile to gain a comprehensive understanding of the changes in protein expression levels during stress and recovery stages. In the long term we aim to combine candidate gene analysis with the QTL mapping effort.
Soybean protein profiling
Another major ongoing research program on drought is the NSF funded project: functional genomics of root growth and root signaling in maize under drought stress. As part of the umbrella of plant stress research we are elucidating gene regulatory network of soybean seed development under drought stress using transcriptome, proteome and metabolome tools. We are engineering soybean for the production of drought tolerant plants by translational genomic approach and cloning and over expression or silencing of soybean genes based on transcript and protein profiling studies.
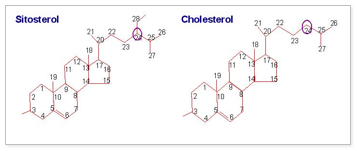
Nutritional genomics: Another major initiative in lab is to identify novel soybean metabolites for human health and nutrition which eventually contribute to improve soybean nutritional value. In collaboration with Prof. Davis Nes at Texas Tech University, we are working to modify the sterol composition of cell membranes to affect drought and temperature tolerance in plants, to modify phytosterol composition to generate resistance to phytopathogens, and to increase the amount of phytosterols in lipid esters of seed oils to benefit human health. Phytosterols in human diet can lower serum cholesterol.
Vuong TD, Sleper DA, Shannon JG, and Nguyen HT. Novel quantitative trait loci for broad-based resistance to soybean cyst nematode (Heterodera glycines Ichinohe) in soybean PI 567516C. Theoretical and Applied Genetics 2010;121(7):1253-1266.
Wu X, Ren C, Joshi T, Vuong T, Xu D, and Nguyen HT. SNP discovery by high-throughput sequencing in soybean. BMC Genomics 2010;11(1):art. no. 469, .
Gutierrez-Gonzalez JJ, Wu X, Gillman JD, Lee JD, Zhong R, Yu O, Shannon G, Ellersieck M, Nguyen HT, and Sleper DA. Intricate environment-modulated genetic networks control isoflavone accumulation in soybean seeds. BMC Plant Biology 2010;10, art. no. 105
Lee JD, Shannon JG, Vuong TD, Moon H, Nguyen HT, Tsukamoto C and Chung G. Genetic diversity in wild soybean (Glycine soja Sieb. and Zucc.) accessions from southern islands of Korean peninsula. Plant Breeding 2010;129(3):257-263.
Manavalan LP, Guttikonda SK, Nguyen VT, Shannon JG and Nguyen HT. Evaluation of diverse soybean germplasm for root growth and architecture. Plant and Soil 2010;330(1):503-514.
Neelakandan AK, Nguyen HTM, Kumar R, Tran LSP, Guttikonda SK, Quach TN, Aldrich DL, Nes WD and Nguyen HT. Molecular characterization and functional analysis of Glycine max sterol methyl transferase 2 genes involved in plant membrane sterol biosynthesis. Plant Molecular Biology 2010;74(4):503-518.
Pham TA, Hill CB, Miles MR, Nguyen BT, Vu TT, Vuong TD, VanToai TT, Nguyen HT and Hartman GL. Evaluation of soybean for resistance to soybean rust in Vietnam. Field Crops Research 2010;117(1):131-138.
Vuong TD, Sleper DA, Shannon JG and Nguyen HT. Novel quantitative trait loci for broad-based resistance to soybean cyst nematode (Heterodera glycines Ichinohe) in soybean PI 567516C. Theoretical and Applied Genetics 2010;121(7):1253-1266.
Wang Z, Libault M, Joshi T, Valliyodan B, Nguyen H, Xu D, Stacey G and Cheng J. SoyDB: A knowledge database of soybean transcription factors. BMC Plant Biology 2010;14.
Wu X, Ren C, Joshi T, Vuong T, Xu D and Nguyen HT. SNP discovery by high-throughput sequencing in soybean. BMC Genomics 2010;11(1):
Yamaguchi M, Valliyodan B, Zhang J, Lenoble ME, Yu O, Rogers EE, Nguyen HT and Sharp RE. Regulation of growth response to water stress in the soybean primary root. I. Proteomic analysis reveals region-specific regulation of phenylpropanoid metabolism and control of free iron in the elongation zone. Plant, Cell and Environment 2010;33(2):223-243.
Gutierrez-Gonzalez JJ, Guttikonda SK, Tran LSP, Aldrich DL, Zhong R, Yu O, Nguyen HT, Sleper DA. Differential expression of isoflavone biosynthetic genes in soybean during water deficits. Plant and Cell Physiology 2010;51(6):936-948.
Gutierrez-Gonzalez JJ, Wu X, Gillman JD, Lee J-D, Zhong R, Yu O, Shannon G, Ellersieck M, Nguyen HT, Sleper DA. Intricate environment-modulated genetic networks control isoflavone accumulation in soybean seeds. BMC Plant Biology 2010:105 (in press).
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC and Jackson SA. Genome sequence of the palaeopolyploid soybean. Nature 2010;463(7278):178-183.
Libault M, Joshi T, Takahashi K, Hurley-Sommer A, Puricelli K, Blake S, Finger RE, Taylor CG, Xu D, Nguyen HT, Stacey G. Plant Physiology 2009;151(3):1207-1220.
Gutierrez-Gonzalez JJ, Wu X, Zhang J, Lee J-D, Ellersieck M, Shannon JG, Yu O, Nguyen HT, Sleper DA. Genetic control of soybean seed isoflavone content: Importance of statistical model and epistasis in complex traits. Theoretical and Applied Genetics 2009;119:1069-1083.
Rogers EE, Wu X, Stacey G, Nguyen HT. Two MATE proteins play a role in iron efficiency in soybean. Journal of Plant Physiology 2009;166:1453-1459.
Manalavan LP, Guttikonda SK, Phan Tran, LS, Nguyen HT. Physiological and molecular approaches to improve drought resistance in soybean. Pland and Cell Physiology 2009;50:1260-1276.
Poroyko V, Spollen WG, Hejlek LG, Hernandez AG, LeNoble ME, Davis G, Nguyen HT, Springer GK, Sharp RE, and Bohnert HJ. Comparing regional transcript profiles from maize primary roots under well-watered and low water potential conditions. J. Exp. Bot. 2007;58:279-290.
Harris K, Subudhi PK, Borrell A, Jordan D, Rosenow D, Nguyen HT , Klein P, Klein R, and Mullet J. Sorghum stay-green QTL individually reduce post-flowering drought-induced leaf senescence. J. Exp. Bot. 2007;58:327-338.
Li Y, Uhm T, Ren C, Wu C, Santos TS, Lee MK, Yan B, Santos F, Zhang A, Scheuring C, Sanchez A, Millena AC, Nguyen HT, Kou H, Liu D, Zhang HB. A plant-transformation-competent BIBAC/BAC-based map of rice for functional analysis and genetic engineering of its genomic sequence. Genome 2007;50:278-288.
Kumar R, Qiu J, Joshi T, Valliyodan B, Xu D, Nguyen HT. Single Feature Polymorphism Discovery in Rice. PLoS ONE 2007;2(3):e284.
Shannon JG, Wrather JA, Sleper DA, Robbins RT, Nguyen HT, and Anand S. Registration of Jake soybean. J. of Crop Registrations 2007;1:29-30.
Shannon JG, Wrather JA, Sleper DA, Nguyen HT, and Anand SC. Registration of Stoddard soybean. J. of Crop Registrations 2007;1:28-29.
Zhang XC, Wu X, Findley S, Wan J, Libault M, Nguyen HT, Cannon SB, Stacey G. Molecular evolution of lysin motif-type receptor-like kinases in plants. Plant Physiol. 2007;144(2):623-36.
Zhu J, Alvarez S, Marsh E, LeNoble ME, Cho IJ, Sivaguru M, Chen S, Nguyen HT, Wu Y, Schachtman DP, Sharp RE. Cell wall proteome in the maize primary root elongation zone. II. Region-specific changes in water soluble and lightly ionically bound proteins under water deficit. Plant Physiol. 2007;145:1533-1548.
Lee JD, Yu JK, Hwang YH, Blake S, So YS, Lee GJ, Nguyen HT, and Shannon JG. Genetic diversity of wild soybean (Glycine soja Sieb. & Zucc) accessions from South Korea and other countries. Crop Science 2008;48:606-616.
Wu X, Zhong G, Findley S, Cregan P, Stacey G, and Nguyen HT. Genetic marker anchoring by six-dimensional pools for development of a soybean physical map. BMC Genomics 2008;9:art. no. 28.
Shoemaker RC, Grant D, Olson T, Warren WC, Wing R, Yu Y, Kim H, Cregan P, Joseph B, Futrell-Griggs M, Nelson W, Davito J, Walker J, Wallis J, Kremitski C, Scheer D, Clifton SW, Graves T, Nguyen HT, Wu X, Luo M, Dvorak J, Nelson R, Cannon S, Tomkins J, Schmutz J, Stacey G, and Jackson S. Microsatellite discovery from BAC end sequences and genetic mapping to anchor the soybean physical and genetic maps. Genome 2008;51:294-302.
Spollen WG, Tao W, Valliyodan B, Chen K, Hejlek LG, Kim JJ, LeNoble ME, Zhu J, Bohnert HJ, Henderson D, Schachtman DP, Davis GE, Springer GK, Sharp RE and Nguyen HT. Spatial distribution of transcript changes in maize primary root elongation zone at low water potential. BMC Plant Biology 2008;8:art. no. 32.
Flores T, Karpova O, Su X, Zeng P, Bilyeu K, Sleper DA, Nguyen HT, and Zhang ZJ. Silencing of GmFAD3 gene by siRNA leads to low alpha-linolenic acids (18:3) of fad3-mutant phenotype in soybean [Glycine max (Merr.)]. Transgenic Research 2008;17:839-850.
Valliyodan B and Nguyen HT. Understanding regulatory networks and engineering for enhanced drought tolerance in plants. Current Opinion in Plant Biology 2006;9:1-7.
Ramalingam J, Pathan MS, Feril O, Miftahudin, Ross K, Ma XF, Mahmoud AA, Layton J, Rodriguez-Milla MA, Chikmawati T, Valliyoda B, Skinner R, Matthews DE, Gustafson JP, and Nguyen HT Structural and functional analyses of the wheat genomes based on expressed sequence tags (ESTs) related to abiotic stresses. Genome 2006;49:1-17.
Nunberg A, Bedell JA, Budiman MA, Citek RW, Clifton SW, Fulton L, Pape D, Cai Z, Joshi T, Nguyen HT, Xu D, and Stacey G. Survey sequencing of soybean elucidates the genome structure and composition. Functional Plant Biology 2006;33:1-9.
Lu P, Shannon JG, Sleper DA, Nguyen HT, Cianzio SR, and Arelli PR. Genetics of Cyst Nematode Resistance in Soybean PIs 467312 and 507354. Euphytica 2006;149:259-265.
Guo B, Sleper DA, Sun J, Nguyen HT, Arelli PR, and Shannon JG. Pooled analysis of data from multiple quantitative trait locus mapping population: QTLs associated with resistance to soybean cyst nematode. Theoretical and Applied Genetics 2006;113:39-48.
Guo B, Sleper DA, Nguyen HT, Arelli PR, and Shannon JG. Quantitative trait loci underlying resistance to three soybean cyst nematode populations in Soybean PI404198A. Crop Science 2006;46:224-233.
Guo B, Sleper DA, Lu P, Shannon JG, Nguyen HT, and Arelli PR. QTLs Associated with Resistance to SCN: Meta analysis of QTL locations. Crop Science 2006;46: 595-602.
Chappel AS, Scaboo AM, Wu X, Nguyen HT, Pantalone VR, and Bilyeu KD. Characterization of the MIPS gene family in Glycine max. Plant Breeding 2006;125:1-8.
Chao S, Lazo GR, You F, et. al. Use of a large-scale Triticeae EST resource to reveal gene expression profiles in hexaploid wheat (Triticum aestivum L.). Genome 2006;49:531-544.
Hazen SP, Pathan MS, Sanchez A, Baxter I, Dunn M, Estes B, Chang HS, Zhu T, Kreps J, and Nguyen HT. Expression profiling of rice segregating for drought tolerance QTLs using a rice genome array. Integr. Funct. Genomics 2005;5: 104-116.
Sharp, RE, V Poroyko, LG Hejlek, WG Spollen, GK Springer, HJ Bohnert, and HT Nguyen. Root growth maintenance during water deficits: physiology to functional genomics. J. Exp. Bot. 2004;55:2343-2351 .
Zhang D, DW Choi, S Wanamaker, RD Fenton, A Chin, M Malatrasi, Y Turuspekov, H Walia, ED Akhunov, P Kianian, C Otto, K Simons, KR Deal, V Echenique, B Stamova, K Ross, GE Butler, L Strader, SD Verhey, R Johnson, S Altenbach, K Kothari, C Tanaka, MM Shah, D Laudencia-Chingcuanco, P Han, RE Miller, CC Crossman, S Chao, GR Lazo, N Klueva, JP Gustafson, SF Kianian, J Dubcovsky, MK Walker-Simmons, KS Gill, J Dvořák, OD Anderson, ME Sorrells, PE McGuire, CO Qualset, HT Nguyen, TJ Close. Construction and evaluation of cDNA libraries for large-scale EST sequencing in wheat (Triticum aestivum L.). Genetics 2004;168:595-608.
Conley EJ, V Nduati, JL Gonzalez-Hernandez, A Mesfin, M Trudeau-Spanjers, S Chao, GR Lazo, DD Hummel, OD Anderson, LL Qi, BS Gill, B Echalier, AM Linkiewicz, J Dubcovsky, ED Akhunov, J Dvořák, JH Peng, NLV Lapitan, MS Pathan, HT Nguyen, XF Ma, Miftahudin, JP Gustafson, RA Greene, ME Sorrells, KG Hossain, V Kalavacharla, SF Kianian, D Sidhu, M Dilbirligi, KS Gill, DW Choi, RD Fenton, TJ Close, PE McGuire, CO Qualset, and JA Anderson. A 2600-locus chromosome bin map of wheat homoeologous group 2 reveals interstitial gene-rich islands and colinearity with rice. Genetics 2004;168:625-637.
Randhawa HS, M Dilbirligi, D Sidhu, M Erayman, D Sandhu, S Bondareva, S Chao, GR Lazo, OD Anderson, Miftahudin, JP Gustafson, B Echalier, LL Qi, BS Gill, ED Akhunov, J Dvořák, AM Linkiewicz, A Ratnasiri, J Dubcovsky, CE Bermudez-Kandianis, RA Greene, ME Sorrells, EJ Wennerlind, JA Anderson, JH Peng, NLV Lapitan, KG Hossain, V Kalavacharla, SF Kianian, MS Pathan, HT Nguyen, TR Endo, TJ Close, PE McGuire, CO Qualset, and KS Gill. Deletion mapping of homoeologous group 6-specific wheat ESTs. Genetics 2004;168:677-686.
Qi LL, B Echalier, S Chao, GR Lazo, GE Butler, OD Anderson, ED Akhunov, J Dvořák, AM Linkiewicz, A Ratnasiri, J Dubcovsky, CE Bermudez-Kandianis, RA Greene, R Kantety, C.M La Rota, JD Munkvold, SF Sorrells, ME Sorrells, M Dilbirligi, D Sidhu, M Erayman, HS Randhawa, D Sandhu, SN Bondareva, KS Gill, AA Mahmoud, XF Ma, Miftahudin, JP Gustafson, EJ Wennerlind, V Nduati, JL Gonzalez-Hernandez, JA Anderson, JH Peng, NLV Lapitan, KG Hossain, V Kalavacharla, SF Kianian, MS Pathan, DS Zhang, HT Nguyen, DW Choi, TJ Close, PE McGuire, CO Qualset, and BS Gill. A chromosome bin map of 16,000 EST loci and distribution of genes among the three genomes of polyploid wheat. Genetics 2004;168:701-712.
Peng JH, H Zadeh, GR Lazo, JP Gustafson, S Chao, OD Anderson, LL Qi, B Echalier, BS Gill, M Dilbirligi, D Sandhu, KS Gill, RA Greene, ME Sorrells, ED Akhunov, J Dvořák, AM Linkiewicz, J Dubcovsky, KG Hossain, V Kalavacharla, SF Kianian, AA Mahmoud, Miftahudin, EJ Wennerlind, JA Anderson, MS Pathan , HT Nguyen, PE McGuire, CO Qualset, and NLV Lapitan. Chromosome bin map of expressed sequence tags in homoeologous group 1 of hexaploid wheat and homoeology with rice and Arabidopsis. Genetics 2004;168:609-623.
Munkvold JD, RA Greene, CE Bermudez-Kandianis, CM La Rota, H Edwards, SF Sorrells,T Dake, D Benscher, R Kantety, AM Linkiewicz, J Dubcovsky, ED Akhunov, J Dvořák, Miftahudin, JP Gustafson, MS Pathan, HT Nguyen, DE Matthews, S Chao, GR Lazo, DD Hummel, OD Anderson, JA Anderson, JL Gonzalez-Hernandez, JH Peng, N Lapitan, LL Qi, B Echalier, BS Gill, KG Hossain, V Kalavacharla, SF Kianian, D Sandhu, M Erayman, KS Gill, PE McGuire, CO Qualset, and ME Sorrells. Group 3 chromosome bin maps of wheat and their relationship to rice chromosome 1. Genetics 2004;168:639-650.
Miftahudin, K Ross, XF Ma, AA Mahmoud, J Layton, M Rodriguez, T Chikmawati, J Ramalingam, O Feril, MS Pathan, G Surlan Momirovic, S Kim, K Chema, P Fang, L Haule, H Struxness, J Birkes, C Yaghoubian, R Skinner, J McAllister, V Nguyen, LL Qi, BS Gill, AM Linkiewicz, J Dubcovsky, ED Akhunov, J Dvořák, M Dilbirligi, KS Gill, JH Peng, NLV Lapitan, CE Bermudez-Kandianis, ME Sorrells, KG Hossain, V Kalavacharla, SF Kianian, GR Lazo, S Chao, OD Anderson, J Gonzalez-Hernandez, EJ Wennerlind, JA Anderson, DW Choi, RD Fenton, TJ Close, PE McGuire, CO Qualset, HT Nguyen, and P Gustafson. Analysis of EST loci on wheat chromosome group 4. Genetics 2004;168:651-663.
Linkiewicz AM, LL Qi, BS Gill, A Ratnasiri, B Echalier, S Chao, GR Lazo, DD Hummel, OD Anderson, ED Akhunov, J Dvořák, MS Pathan, HT Nguyen, JH Peng, NLV Lapitan, Miftahudin, JP Gustafson, CM La Rota, ME Sorrells, KG Hossain, V Kalavacharla, SF Kianian, D Sandhu, SN Bondareva, KS Gill, EJ Wennerlind, JA Anderson, RD Fenton, TJ Close, PE McGuire, CO Qualset, and J Dubcovsky. A 2500-locus bin map of wheat homoeologous group 5 provides new insights on gene distribution and colinearity with rice. Genetics 2004;168:665-676.
Lazo GR, S Chao, DD Hummel, H Edwards, CC Crossman, N Lui, DE Matthews, VL Carollo, DL Hane, FM You, GE Butler, RE Miller, TJ Close, JH Peng, NLV Lapitan, JP Gustafson, LL Qi, B Echalier, BS Gill, M Dilbirligi, D Sandhu, KS Gill, RA Greene, ME Sorrells, ED Akhunov, J Dvořák, AM Linkiewicz, J Dubcovsky, KG Hossain, V Kalavacharla, SF Kianian, AA Mahmoud, Miftahudin, XF Ma, EJ Wennerlind, JA Anderson, MS Pathan , HT Nguyen, PE McGuire, CO Qualset, and OD Anderson. Development of an expressed sequence tag (EST) resource for wheat (Triticum aestivum L.): EST generation, unigene analysis, probe selection, and bioinformatics for a 16,000-locus bin-delineated map. Genetics 2004;168:585-593.
Hossain KG, V Kalavacharla, GR Lazo, J Hegstad, MJ Wentz, K Simons, S Gehlhar, JL Rust, RR Syamala, K Obeori, B Suresh, P Karunadharma, S Chao, OD Anderson, LL Qi, B Echalier, BS Gill, AM Linkiewicz, A Ratnasiri, J Dubcovsky, ED Akhunov, J Dvořák, Miftahudin, K Ross, JP Gustafson, D Sidhu, M Dilbirligi, KS Gill, JH Peng, NLV Lapitan, RA Greene, CE Bermudez-Kandianis, ME Sorrells, O Feril, MS Pathan, HT Nguyen, JL Gonzalez-Hernandez, EJ Wennerlind, JA Anderson, D Fenton, TJ Close, PE McGuire, CO Qualset, and SF Kianian. A chromosome bin map of 2148 EST loci of wheat homoeologous group 7. Genetics 2004;168:687-699.
Nguyen TT, Klueva N, Chamareck V, Aarti A, Magpantay G, Millena AC, Pathan MS, and Nguyen HT. Saturation mapping of QTL regions and identification of putative candidate genes for drought tolerance in rice. Mol. Gen. Genet. 2004;272:35-46.
Babu RC, J Zhang, A Blum, THD Ho, R Wu, and HT Nguyen. HVA1, a LEA gene from barley confers dehydration tolerance in transgenic rice (Oryza sativa L.) via cell membrane protection. Plant Science 2004; 166:855-862.
Robin S, MS Pathan, B Courtois, R Lafitte, C Scarandang, S Lanceras, M Amante, HT Nguyen, and Z Li. Mapping osmotic adjustment in an advanced backcross inbred population of rice. Theor. Appl. Genetics 2004;107: 1288-1296
Akhunov ED, AR Akhunova, AM Linkiewicz, J Dubcovsky, D Hummel, G Lazo, S Chao, OD Anderson, J David , LL Qi, B Echalier, BS Gill, Miftahudin, P Gustafson, CM La Rota, ME Sorrells, D Zhang , HT Nguyen, V Kalavacharla, K Hossain, SF Kianian, J Peng, NLV Lapitan, EJ Wennerlind , V Nduati, JA Anderson, D Sidhu, KS Gill, PE McGuire, CO Qualset, and J Dvorak. Synteny perturbations between wheat homoeologous chromosomes caused by locus duplications and deletions correlate with recombination rates along chromosome arm.Proc. Natural Academy of Science 2003;(USA) 100: 10836-10841.
Sorrells ME, CM La Rota, CE Bermudez-Kandianis, RA Greene, R Kantety, JD Munkvold, Miftahudin, A Mahmoud, JP Gustafson, LL Qi, B Echalier, BS Gill, D Matthews, G Lazo, S Chao, OD Anderson, H Edwards, AM Linkiewicz, J Dubcovsky, ED Akhunov, J Dvorak, D Zhang , HT Nguyen, J Peng, NLV Lapitan, JL Gonzalez-Hernandez, JA Anderson, KG Hossain, V Kalavacharla, SF Kianian, DW Choi, TJ Close, M Dilbirligi, KS Gill, C Steber, MK Walker-Simmons, PE McGuire, and CO Qualset. Comparative DNA Sequence Analysis of Wheat and Rice Genomes. Genome Research 2003;13:1818-1827.
Akhunov ED, JA Goodyear, S Geng, LL Qi, B Echalier, BS Gill, G Lazo, S Chao, OD Anderson, AM Linkiewicz, J Dubcovsky, CM La Rota, ME Sorrells, MD Zhang, HT Nguyen, PJ Gustafson, V Kalavacharla, KG Hossain, SF Kianian, J Peng, NLV Lapitan, JL Gonzalez-Hernandez, JA Anderson, DW Choi, TJ Close, M Dilbirligi, KS Gill, MK Walker-Simmons, C Steber, PE McGuire, CO Qualset, and J Dvorak. The organization and rate of evolution of the wheat transcriptome are correlated with recombination rates along chromosome arms. Genome Research 2003;13:753-763.
Babu RC, BD Nguyen, Varapong Chamarerk, P Shanmugasundaram, P Chezhian, P Jeyaprakash, SK Ganesh, A Palchamy, S Sadasivam, S Sarkarung, LJ Wade and HT Nguyen. Genetic analysis of drought resistance in rice by molecular markers: association between secondary traits and field performance. Crop Sciences 2003;43:1457-1469
- Wiese-Fales J. Sequencing of soybean genome adds potential to research (Mar 2010)
- Researchers fight world hunger by mapping the soybean genome (Feb 2010)
- Bean Breakthrough. Illumination (Spring 2010)
- Soybean genome mapping could lead to improved varieties. Mizzou Weekly (Jan 2010)
- Mertens R. A dry subject: CAFNR researchers study the effects of drought by making their own (June 2009)
- Henry Nguyen elected AAAS Fellow (January 2009)
- Building soybean's 'parts list'. Delta Farm Press (Jan 2009)
- Soybean database will help breeders engineer better-performing plants (Dec 2008)
- DOE JGI releases soybean genome assembly to enable worldwide bioenergy research efforts (Jan 2008)
- Monsanto is shifting its focus from corn to new biotech soybeans (Jan 2008)
- Ganey T. MU opens soybean genetics lab. Columbia Daily Tribune (Oct 2, 2006)
- Horstmeier G. Soybean central. Mizzou News (Dec 2003)
- Fellow, American Association for the Advancement of Science, 2009
- Research Award, Missouri Soybean Association, 2005
- Fellow, American Society of Agronomy, 2000
- Fellow, Crop Science Society of America, 1999
- President's Academic Achievement Award, Texas Tech University, 1996
- Outstanding Researcher, College of Agricultural Sciences and Natural Resources, Texas Tech University, 1995
- Young Crop Scientist Award, Crop Science Society of America, 1990
- Presidential Young Investigator Award, National Science Foundation, 1986